

Jul 9, 2019
Big data and artificial intelligence are transforming the current technological landscape. However, these advances rarely make an impact in clinical practice. We address the unique challenges that arise when applying big data and machine learning techniques to healthcare data with the goal of improving patient care and clinical research.
We work closely with clinical departments to develop predictive models and identify risk factors for clinical outcomes at scale.
We are actively involved in addressing some of these broad clinical questions featured below, and pilot projects with additional departments are in the works.
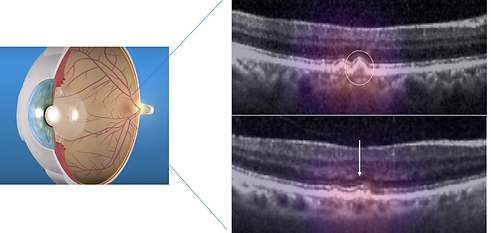
Age-related Macular Degeneration (AMD) is the leading cause of blindness for elderly individuals. The disease is currently irreversible, but the recent widespread adoption of optical coherence tomography (OCT) has made it easier to diagnose. We've developed methods for automatically detecting risk-factors for advanced disease. Using these risk factors and the electronic health record, can we define, characterize, and predict the onset and deterioration of Age-related Macular Degeneration?
Not much needs to be said about the global impact of the COVID-19 pandemic. We developed a machine learning based proxy PCR to address a potential testing shortage, made recommendations about discharge protocol for hospitalized patients, and modeled the impact of surges on UCLA health resources. Some of our models were used to guide the system through the pandemic.
Currently, we are trying to identify risk factors for COVID-related complications and breakthrough cases.
There are several central challenges that span all big data applications to the medical domain. We are always thinking about these issues in our projects:
Limited data: Despite nearly all health records being digital now, there is a disconnect between modern big data approaches and the clinical research cycle. Big data approaches often require troves of annotated data, which is often not the case for areas of active clinical research, where the phenomenon itself is still being defined or simply rare, and large-scale annotation is not feasible. We explore methods for overcoming this limitation.
Data heterogeneity: Many different types of data are collected and aggregated as part of the electronic health record. These data can span different modalities (e.g. imaging, text), biological domains, and temporal scale. We are interested in developing pipelines and techniques to ingest, align, and jointly represent these data using modern and traditional data fusion approaches.
Equity and Diversity: We are committed to ensuring our models and findings are as free from bias as possible and leverage the diversity in our data and statistical techniques to do so. We are also always on the lookout for advances that help address bias in computational modeling.
Privacy: It goes without saying that patient privacy is paramount in our work. We are interested in exploring methods for generating insights from data across institutions without the need for sharing raw data.
We are also engaged in translating research into practice. The Technology Core is involved in building the technical infrastructure for new technology to be deployed in the clinic. Students in the technology core are exposed to a wide range of tools and best practices in cloud computing and software development.
Unfortunately, we are not taking new applications for the technology core at this time.
I earned my PhD in Computational Cognition (Psychology) at UCLA, where I also completed my Bachelor's degree in Cognitive Science and Statistics. I've also briefly worked in data science-related roles at Voxelcloud, Delta Brain, and the JPL prior to joining Computational Medicine at UCLA.




Eran Halperin, PhD
Eleazar Eskin, PhD
Srinivas R Sadda, MD
David Goodman-Meza, MD, MAS
OHIA